FAQ
There are a number of ways to explore GenEpiO’s content.
- There are a variety of ontology search engines that provide a convenient way browse the GenEpiO hierarchy and search for terms:
- EMBL-EBI Ontology Lookup Service
- National Center for Biomedical Ontology BioPortal
- He Group (University of Michigan Medical School) OntoBee
- Visit the https://github.com/genepio/genepio/ repository where GenEpiO files reside. We have introduced a number of terms into the main file, **genepio.owl**, which also imports terms from over 25 OBO Foundry family ontologies (in the /imports folder).
- View GenEpiO using the free Protégé OWL ontology editor produced by Stanford University. After downloading the GenEpiO github archive files, and running Protégé, open the “genepio.owl” file and click on the Classes tab. There you can drill-down the entity hierarchy; or use the search box in upper right to find terms.
- Preview our Proof Sheet application we are developing for previewing the data-collection and reporting elements of an ontology.
- Explore the GenEpiO Upper Level Schema spreadsheet which gives an overview of GenEpiO’s scope.
- An introductory slideshow is available.
Note that GenEpiO is under development (many terms have a curation status of “requires discussion”) and its existing content requires a few more editing passes for proper description of terms.
GenEpiO terms/concepts are organized under the upper level Basic Formal Ontology (BFO) and the closely related Ontology for Biomedical Investigations (OBI). Selected BFO terms form the top level of GenEpiO; this example shows the “information content entity” tree that houses many terms:
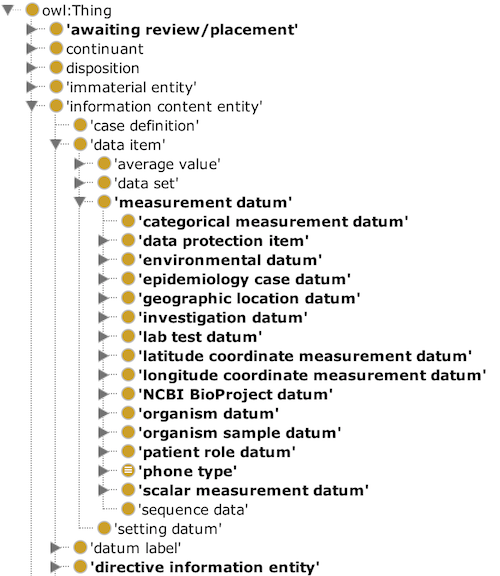

Since the implementation of a new term may require discussion, turn-around times are not fixed. Stakeholders that need a new term can simply email it and its definition in to info (at) genepio.org or to the GenEpiO forum, where it can be routed to the appropriate editor. However, we have a standard web-driven ontology term addition request form we encourage people to use because it contains more detailed instructions that smooth the editorial cycle.
- Go to the GenEpiO issue tracker at https://github.com/GenEpiO/genepio/issues
- Click on “New issue”
- Fill in the “Term Request / Revision / Import Form” template
- This form is drawn from the official OBI specification for term entry; further guidelines for metadata annotation are available here.
A request to integrate GenEpiO with (i.e. import) an existing term from another (preferably OBOFoundry) ontology should be carried out as per the instructions described on the Issues tracker page. Integration is performed according to MIEROT principles of term adoption. (Currently an OWL format import file for each 3rd party ontology’s subset of terms is created manually using the http://ontofox.hegroup.org service which operates on a text file input containing a short list of target term ids. The OntoFox import files are included in GenEpiO github ontofox_input folder; and the generated OWL files are in the /imports/ folder.)
When a draft entry in GenEpiO has been entered, it will be given a “curation status” state described here: http://obi-ontology.org/page/OBI_Minimal_metadata, which is expected to move towards the “ready for release” status as editing is completed. No other formal requirements exist here yet.
Ontology definition owes a great debt to traditional dictionary compilation, with a behind-the-scenes focus on keeping a record of a term’s editor, creation date and source(s). There are a few differences. Rather than state a term’s grammar, one focuses on the (OBO) semantics of a term – whether it refers to a process, or a persistent entity (material or immaterial) or one of its behavioural states, characteristics or observable qualities. Rather than state first known use of a term, one attributes the term or its definition to a source ontology, database or document.
The textual definition of a term is expected to be in an Aristotelian form, which – if it sounds right – breeds subclasses; An A is a B which has a feature or differentia C. “A Salmonella antigenic formula is an antigenic formula that is about a salmonella isolate.” It is expected that the Technical working group will translate this into BFO/OBI logical relations as much as possible.
Some terms may be amenable to successive definition enhancements over time. For example, a term could start off as a relatively simple OBI “measurement datum” which involves recording a medical or scientific observation at some point in time, a “pH measurement” for example. The status can be updated to “ready for release”, but the term could also be further enriched over time with additional logical relations such as “’has measurement unit label’ some pH”, an “’is about’ acidity” relation, a specification of its lower and upper integer measurement bounds (0 – 14 usually), and even the stated implication that any measurement datum is associated with a date/time of observation.